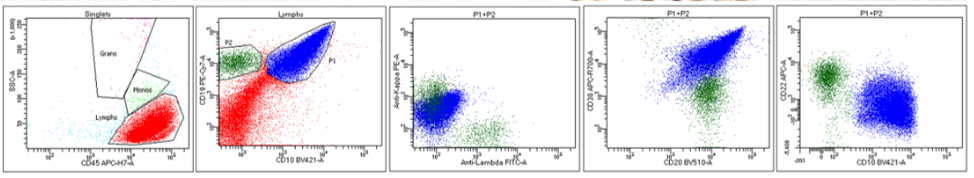
Flow cytometry:
IHC stains:

Cytogenetics:
46,XY,der(11)del(11)(p13p15.1)dup(11)(q22.3q23.3)[11]/46,XY,der(11)del(11)(p13p15.1)trp(11)(q22.3q23.3)[3]/ 46,XY[3]
FISH: negative for MYC rearrangement
Microarray: segmental gains within 11q with adjacent terminal loss. Solid blue bar representing area of gain (green arrow) and amplification (black arrow), and solid red bar indicating area of loss (red arrow) within chromosome 11; the green bracket outlining an area of chromothripsis on 11p (pink arrow).
46,XY,der(11)del(11)(p13p15.1)dup(11)(q22.3q23.3)[11]/46,XY,der(11)del(11)(p13p15.1)trp(11)(q22.3q23.3)[3]/ 46,XY[3]
FISH: negative for MYC rearrangement
Microarray: segmental gains within 11q with adjacent terminal loss. Solid blue bar representing area of gain (green arrow) and amplification (black arrow), and solid red bar indicating area of loss (red arrow) within chromosome 11; the green bracket outlining an area of chromothripsis on 11p (pink arrow).
Diagnosis: Burkitt-like lymphoma with 11q aberration
Burkitt-like lymphoma with 11q aberration (BLL-11q), a new provisional entity in the 2016 WHO classification, comprises cases with morphologic, phenotypic, and gene expression resemblance to BL, but lacking MYC translocation (MYC-) and harboring characteristic proximal 11q gains and distal 11q loss. BLL-11q lymphomas have mutational landscape different from BL. Previous studies defined the typical gained region as 11q23.3, and the lost region as 11q24-qter. Collectively, their results suggest the upregulation of oncogenes PAFAH1B2, USP2, and CBL located in the gained regions of 11q23, and corresponding downregulation of tumor suppressor candidate genes FLI1, ETS1, TBRG1, and EI24 located in the regions of 11q24-qter loss. There are 2 different types of 11q rearrangements seen in BLL-11q cases: a large duplication of 11q comprising more than 50 Mb with terminal deletion, and a small duplication of 11q comprising less than 20 Mb with an additional gain within the duplicated material as well as terminal deletion. BLL-11q cases occur over a wide age range, but are more common in children and young adults. They are more frequently nodal than BL, and tend to present as a single dominant mass or conglomerate mass. Patients tend to present with limited disease without involvement of bone marrow or CSF, and prognosis appears to be favorable, similar to classical BL.
The diagnosis of BLL-11q is based on the presence of characteristic gain/loss patterns of 11q, together with BL/BLL morphology, GCB phenotype, and lack of MYC rearrangement. The characteristic 11q aberration is key to making the diagnosis, but its presence alone is neither specific nor diagnostic since it may also be present in MYC+ BL or DLBCL. The most sensitive modality for detecting this characteristic cytogenetic finding is DNA microarray. The 11q aberration can be visualized by chromosomal analysis, as seen in our two cases. However, chromosomal analysis relies on tumor cell viability and metaphase morphology, and the finding may not be characteristic for this aberration when resolution is low. Another potential diagnostic strategy is FISH for chromosome 11 abnormalities. Commercially available FISH probes for chromosome 11 regions may be used to detect gains within 11q and 11q terminal loss. Some groups have suggested a diagnostic strategy of MYC and chromosome 11 FISH, with reflex to DNA microarray if inconclusive. Due to the variation of gain/loss spots among the cases, depending on the probes used, FISH method alone may miss some cases. On the other hand, some cases may be falsely called positive. Microarray or FISH should be tested on all MYC-negative high grade mature B-cell lymphoma with BL/BLL morphology. Some study showed that the phenotype of BLL-11q was similar to that of BL except for a characteristic lower expression of CD38, and coexpression of CD16/CD56. This characteristic expression profile may be used to screen such cases and determine the need for DNA microarray/FISH for detection of 11q abnormalities.
Ref:
Burkitt-like lymphoma with 11q aberration (BLL-11q), a new provisional entity in the 2016 WHO classification, comprises cases with morphologic, phenotypic, and gene expression resemblance to BL, but lacking MYC translocation (MYC-) and harboring characteristic proximal 11q gains and distal 11q loss. BLL-11q lymphomas have mutational landscape different from BL. Previous studies defined the typical gained region as 11q23.3, and the lost region as 11q24-qter. Collectively, their results suggest the upregulation of oncogenes PAFAH1B2, USP2, and CBL located in the gained regions of 11q23, and corresponding downregulation of tumor suppressor candidate genes FLI1, ETS1, TBRG1, and EI24 located in the regions of 11q24-qter loss. There are 2 different types of 11q rearrangements seen in BLL-11q cases: a large duplication of 11q comprising more than 50 Mb with terminal deletion, and a small duplication of 11q comprising less than 20 Mb with an additional gain within the duplicated material as well as terminal deletion. BLL-11q cases occur over a wide age range, but are more common in children and young adults. They are more frequently nodal than BL, and tend to present as a single dominant mass or conglomerate mass. Patients tend to present with limited disease without involvement of bone marrow or CSF, and prognosis appears to be favorable, similar to classical BL.
The diagnosis of BLL-11q is based on the presence of characteristic gain/loss patterns of 11q, together with BL/BLL morphology, GCB phenotype, and lack of MYC rearrangement. The characteristic 11q aberration is key to making the diagnosis, but its presence alone is neither specific nor diagnostic since it may also be present in MYC+ BL or DLBCL. The most sensitive modality for detecting this characteristic cytogenetic finding is DNA microarray. The 11q aberration can be visualized by chromosomal analysis, as seen in our two cases. However, chromosomal analysis relies on tumor cell viability and metaphase morphology, and the finding may not be characteristic for this aberration when resolution is low. Another potential diagnostic strategy is FISH for chromosome 11 abnormalities. Commercially available FISH probes for chromosome 11 regions may be used to detect gains within 11q and 11q terminal loss. Some groups have suggested a diagnostic strategy of MYC and chromosome 11 FISH, with reflex to DNA microarray if inconclusive. Due to the variation of gain/loss spots among the cases, depending on the probes used, FISH method alone may miss some cases. On the other hand, some cases may be falsely called positive. Microarray or FISH should be tested on all MYC-negative high grade mature B-cell lymphoma with BL/BLL morphology. Some study showed that the phenotype of BLL-11q was similar to that of BL except for a characteristic lower expression of CD38, and coexpression of CD16/CD56. This characteristic expression profile may be used to screen such cases and determine the need for DNA microarray/FISH for detection of 11q abnormalities.
Ref:
- Zhang L, Brown LE, Bowen LM, Application of 2016 WHO classification in the diagnosis of pediatric high grade MYC-negative mature B-cell lymphoma with Burkitt-like morphological features. J Clin Pathol 2020. doi:10.1136/jclinpath-2019-206267